arviz.plot_density#
- arviz.plot_density(data, group='posterior', data_labels=None, var_names=None, filter_vars=None, combine_dims=None, transform=None, hdi_prob=None, point_estimate='auto', colors='cycle', outline=True, hdi_markers='', shade=0.0, bw='default', circular=False, grid=None, figsize=None, textsize=None, labeller=None, ax=None, backend=None, backend_kwargs=None, show=None)[source]#
Generate KDE plots for continuous variables and histograms for discrete ones.
Plots are truncated at their 100*(1-alpha)% highest density intervals. Plots are grouped per variable and colors assigned to models.
- Parameters:
- data
InferenceDataor iterable ofInferenceData Any object that can be converted to an
arviz.InferenceDataobject, or an Iterator returning a sequence of such objects. Refer to documentation ofarviz.convert_to_dataset()for details.- group{“posterior”, “prior”}, default “posterior”
Specifies which InferenceData group should be plotted. If “posterior”, then the values in
posterior_predictivegroup are compared to the ones inobserved_data, if “prior” then the same comparison happens, but with the values inprior_predictivegroup.- data_labels
listofstr, defaultNone List with names for the datasets passed as “data.” Useful when plotting more than one dataset. Must be the same shape as the data parameter.
- var_names
listofstr, optional List of variables to plot. If multiple datasets are supplied and
var_namesis not None, will print the same set of variables for each dataset. Defaults to None, which results in all the variables being plotted.- filter_vars{
None, “like”, “regex”}, defaultNone If
None(default), interpretvar_namesas the real variables names. If “like”, interpretvar_namesas substrings of the real variables names. If “regex”, interpretvar_namesas regular expressions on the real variables names. See this section for usage examples.- combine_dims
set_likeofstr, optional List of dimensions to reduce. Defaults to reducing only the “chain” and “draw” dimensions. See this section for usage examples.
- transform
callable() Function to transform data (defaults to
Nonei.e. the identity function).- hdi_prob
float, default 0.94 Probability for the highest density interval. Should be in the interval (0, 1]. See this section for usage examples.
- point_estimate
str, optional Plot point estimate per variable. Values should be ‘mean’, ‘median’, ‘mode’ or None. Defaults to ‘auto’ i.e. it falls back to default set in
rcParams.- colors
strorlistofstr, optional List with valid matplotlib colors, one color per model. Alternative a string can be passed. If the string is
cycle, it will automatically choose a color per model from matplotlib’s cycle. If a single color is passed, e.g. ‘k’, ‘C2’ or ‘red’ this color will be used for all models. Defaults tocycle.- outlinebool, default
True Use a line to draw KDEs and histograms.
- hdi_markers
str A valid
matplotlib.markerslike ‘v’, used to indicate the limits of the highest density interval. Defaults to empty string (no marker).- shade
float, default 0 Alpha blending value for the shaded area under the curve, between 0 (no shade) and 1 (opaque).
- bw
floatorstr, optional If numeric, indicates the bandwidth and must be positive. If str, indicates the method to estimate the bandwidth and must be one of “scott”, “silverman”, “isj” or “experimental” when
circularis False and “taylor” (for now) whencircularis True. Defaults to “default” which means “experimental” when variable is not circular and “taylor” when it is.- circularbool, default
False If True, it interprets the values passed are from a circular variable measured in radians and a circular KDE is used. Only valid for 1D KDE.
- grid
tuple, optional Number of rows and columns. Defaults to
None, the rows and columns are automatically inferred. See this section for usage examples.- figsize(
float,float), optional Figure size. If
Noneit will be defined automatically.- textsize
float, optional Text size scaling factor for labels, titles and lines. If
Noneit will be autoscaled based onfigsize.- labellerLabeller, optional
Class providing the method
make_label_vertto generate the labels in the plot titles. Read the Label guide for more details and usage examples.- ax2D array_like of
matplotlib AxesorBokeh Figure, optional A 2D array of locations into which to plot the densities. If not supplied, ArviZ will create its own array of plot areas (and return it).
- backend{“matplotlib”, “bokeh”}, default “matplotlib”
Select plotting backend.
- backend_kwargs
dict, optional These are kwargs specific to the backend being used, passed to
matplotlib.pyplot.subplots()orbokeh.plotting.figure. For additional documentation check the plotting method of the backend.- showbool, optional
Call backend show function.
- data
- Returns:
- axes2D
ndarrayofmatplotlib AxesorBokeh Figure
- axes2D
See also
plot_distPlot distribution as histogram or kernel density estimates.
plot_posteriorPlot Posterior densities in the style of John K. Kruschke’s book.
Examples
Plot default density plot
>>> import arviz as az >>> centered = az.load_arviz_data('centered_eight') >>> non_centered = az.load_arviz_data('non_centered_eight') >>> az.plot_density([centered, non_centered])
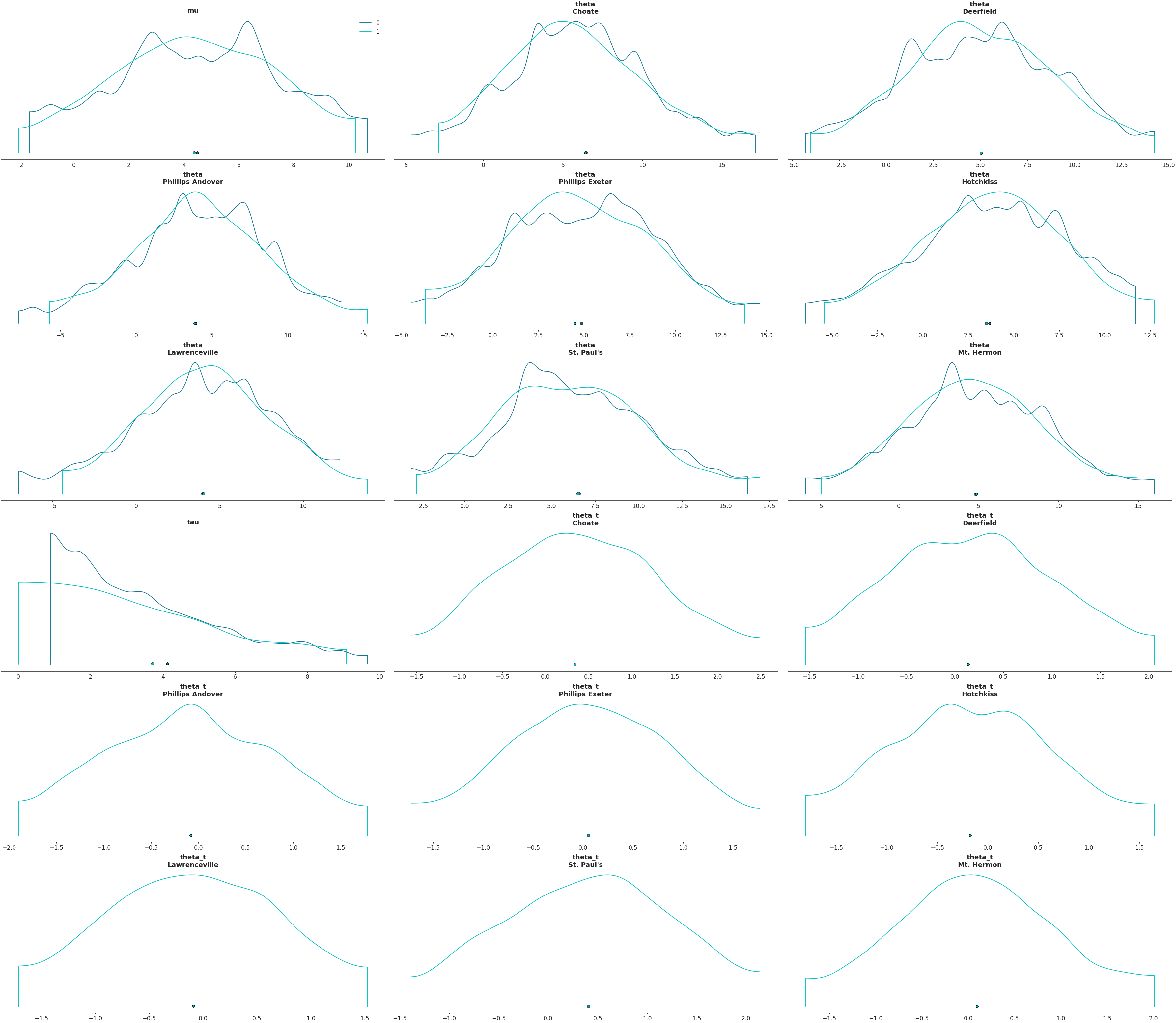
Plot variables in a 4x5 grid
>>> az.plot_density([centered, non_centered], grid=(4, 5))
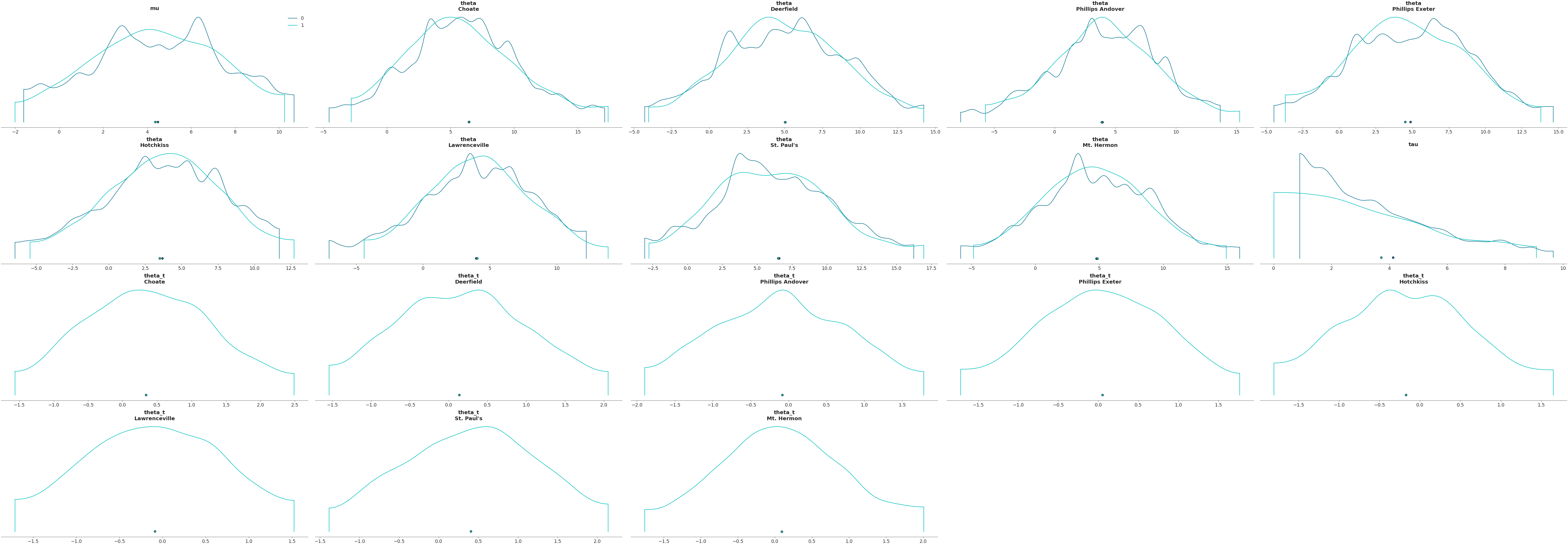
Plot subset variables by specifying variable name exactly
>>> az.plot_density([centered, non_centered], var_names=["mu"])
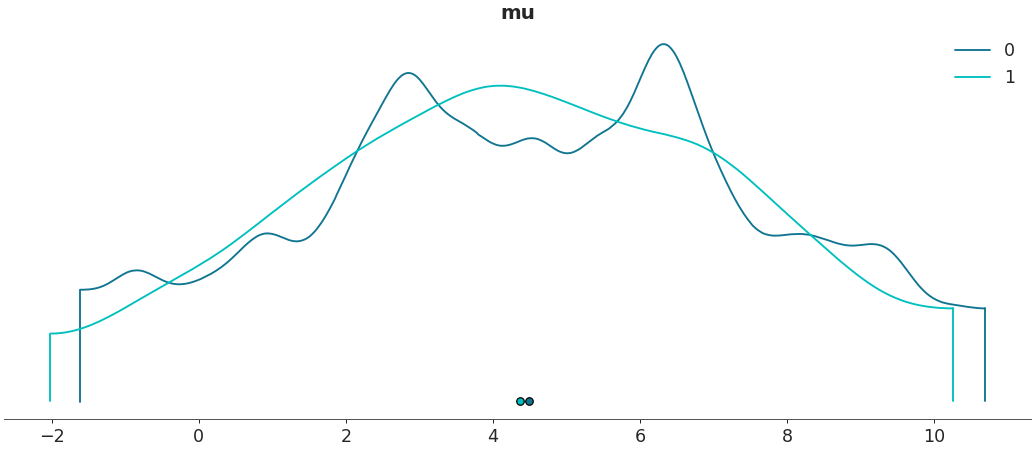
Plot a specific
az.InferenceDatagroup>>> az.plot_density([centered, non_centered], var_names=["mu"], group="prior")
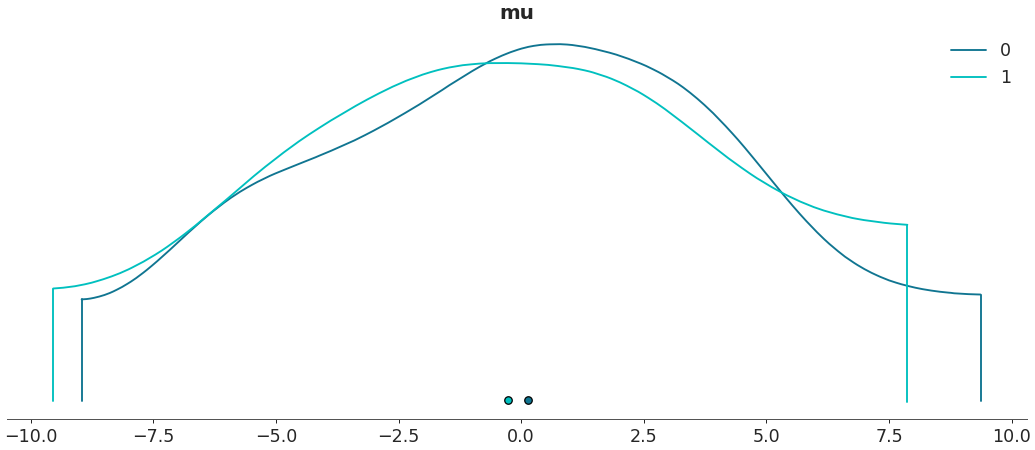
Specify highest density interval
>>> az.plot_density([centered, non_centered], var_names=["mu"], hdi_prob=.5)
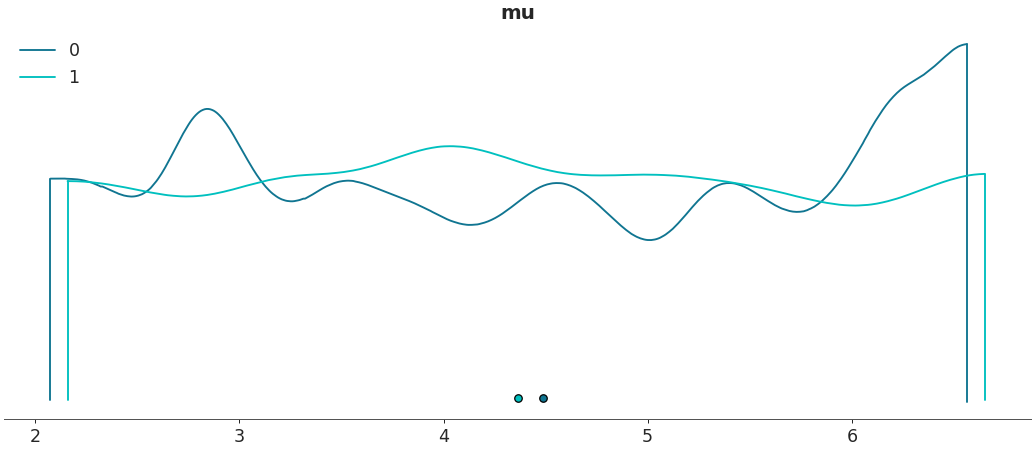
Shade plots and/or remove outlines
>>> az.plot_density([centered, non_centered], var_names=["mu"], outline=False, shade=.8)
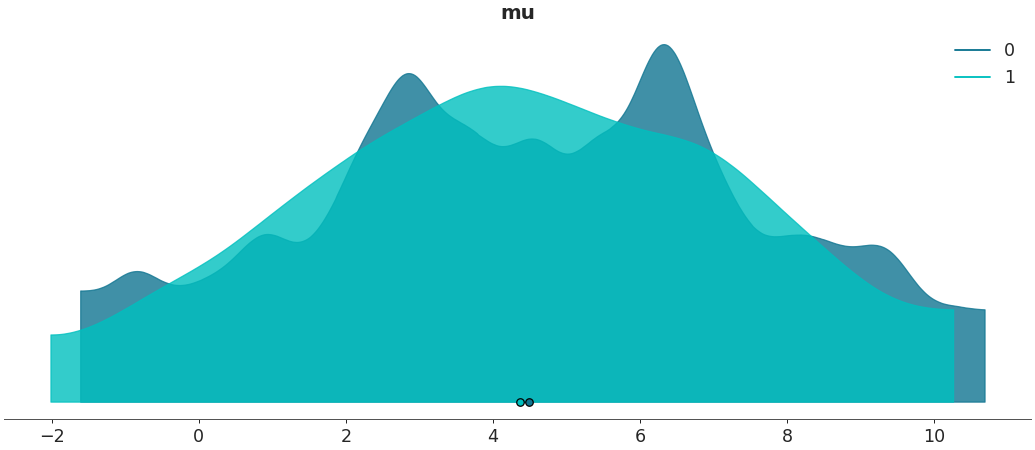
Specify binwidth for kernel density estimation
>>> az.plot_density([centered, non_centered], var_names=["mu"], bw=.9)