arviz.plot_ppc#
- arviz.plot_ppc(data, kind='kde', alpha=None, mean=True, observed=True, color=None, colors=None, grid=None, figsize=None, textsize=None, data_pairs=None, var_names=None, filter_vars=None, coords=None, flatten=None, flatten_pp=None, num_pp_samples=None, random_seed=None, jitter=None, animated=False, animation_kwargs=None, legend=True, labeller=None, ax=None, backend=None, backend_kwargs=None, group='posterior', show=None)[source]#
Plot for posterior/prior predictive checks.
- Parameters
- data: az.InferenceData object
arviz.InferenceDataobject containing the observed and posterior/prior predictive data.- kind: str
Type of plot to display (“kde”, “cumulative”, or “scatter”). Defaults to kde.
- alpha: float
Opacity of posterior/prior predictive density curves. Defaults to 0.2 for
kind = kdeand cumulative, for scatter defaults to 0.7.- mean: bool
Whether or not to plot the mean posterior/prior predictive distribution. Defaults to
True.- observed: bool, default True
Whether or not to plot the observed data.
- color: str
Valid matplotlib
color. Defaults toC0.- color: list
List with valid matplotlib colors corresponding to the posterior/prior predictive distribution, observed data and mean of the posterior/prior predictive distribution. Defaults to [“C0”, “k”, “C1”].
- gridtuple
Number of rows and columns. Defaults to None, the rows and columns are automatically inferred.
- figsize: tuple
Figure size. If None, it will be defined automatically.
- textsize: float
Text size scaling factor for labels, titles and lines. If None, it will be autoscaled based on
figsize.- data_pairs: dict
Dictionary containing relations between observed data and posterior/prior predictive data. Dictionary structure:
key = data var_name
value = posterior/prior predictive var_name
For example,
data_pairs = {'y' : 'y_hat'}If None, it will assume that the observed data and the posterior/prior predictive data have the same variable name.- var_names: list of variable names
Variables to be plotted, if None all variable are plotted. Prefix the variables by
~when you want to exclude them from the plot.- filter_vars: {None, “like”, “regex”}, optional, default=None
If None (default), interpret var_names as the real variables names. If “like”, interpret var_names as substrings of the real variables names. If “regex”, interpret var_names as regular expressions on the real variables names. A la
pandas.filter.- coords: dict
Dictionary mapping dimensions to selected coordinates to be plotted. Dimensions without a mapping specified will include all coordinates for that dimension. Defaults to including all coordinates for all dimensions if None.
- flatten: list
List of dimensions to flatten in
observed_data. Only flattens across the coordinates specified in thecoordsargument. Defaults to flattening all of the dimensions.- flatten_pp: list
List of dimensions to flatten in posterior_predictive/prior_predictive. Only flattens across the coordinates specified in the
coordsargument. Defaults to flattening all of the dimensions. Dimensions should match flatten excluding dimensions fordata_pairsparameters. Ifflattenis defined andflatten_ppis None, thenflatten_pp = flatten.- num_pp_samples: int
The number of posterior/prior predictive samples to plot. For
kind= ‘scatter’ andanimation = Falseif defaults to a maximum of 5 samples and will set jitter to 0.7. unless defined. Otherwise it defaults to all provided samples.- random_seed: int
Random number generator seed passed to
numpy.random.seedto allow reproducibility of the plot. By default, no seed will be provided and the plot will change each call if a random sample is specified bynum_pp_samples.- jitter: float
If
kindis “scatter”, jitter will add random uniform noise to the height of the ppc samples and observed data. By default 0.- animated: bool
Create an animation of one posterior/prior predictive sample per frame. Defaults to
False. Only works with matploblib backend. To run animations inside a notebook you have to use the nbAgg matplotlib’s backend. Try with %matplotlib notebook or %matplotlib nbAgg. You can switch back to the default matplotlib’s backend with %matplotlib inline or %matplotlib auto. If switching back and forth between matplotlib’s backend, you may need to run twice the cell with the animation. If you experience problems rendering the animation try setting animation_kwargs({‘blit’:False}) or changing the matplotlib’s backend (e.g. to TkAgg) If you run the animation from a script write ax, ani = az.plot_ppc(.)- animation_kwargsdict
Keywords passed to
matplotlib.animation.FuncAnimation. Ignored with matplotlib backend.- legendbool
Add legend to figure. By default
True.- labellerlabeller instance, optional
Class providing the method
make_pp_labelto generate the labels in the plot titles. Read the Label guide for more details and usage examples.- ax: numpy array-like of matplotlib axes or bokeh figures, optional
A 2D array of locations into which to plot the densities. If not supplied, Arviz will create its own array of plot areas (and return it).
- backend: str, optional
Select plotting backend {“matplotlib”,”bokeh”}. Default to “matplotlib”.
- backend_kwargs: bool, optional
These are kwargs specific to the backend being used, passed to
matplotlib.pyplot.subplots()orbokeh.plotting.figure(). For additional documentation check the plotting method of the backend.- group: {“prior”, “posterior”}, optional
Specifies which InferenceData group should be plotted. Defaults to ‘posterior’. Other value can be ‘prior’.
- show: bool, optional
Call backend show function.
- Returns
- axes: matplotlib axes or bokeh figures
See also
Examples
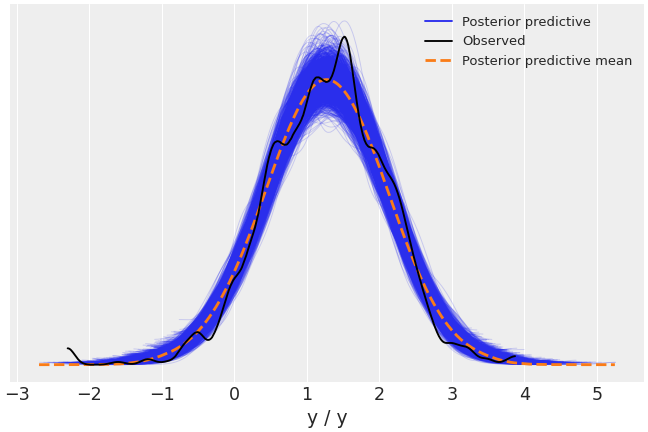
Plot the observed data KDE overlaid on posterior predictive KDEs.
>>> import arviz as az >>> data = az.load_arviz_data('radon') >>> az.plot_ppc(data, data_pairs={"y":"y"})
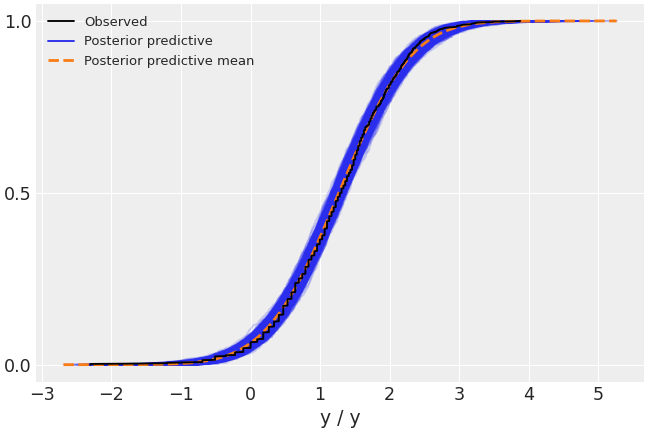
Plot the overlay with empirical CDFs.
>>> az.plot_ppc(data, kind='cumulative')
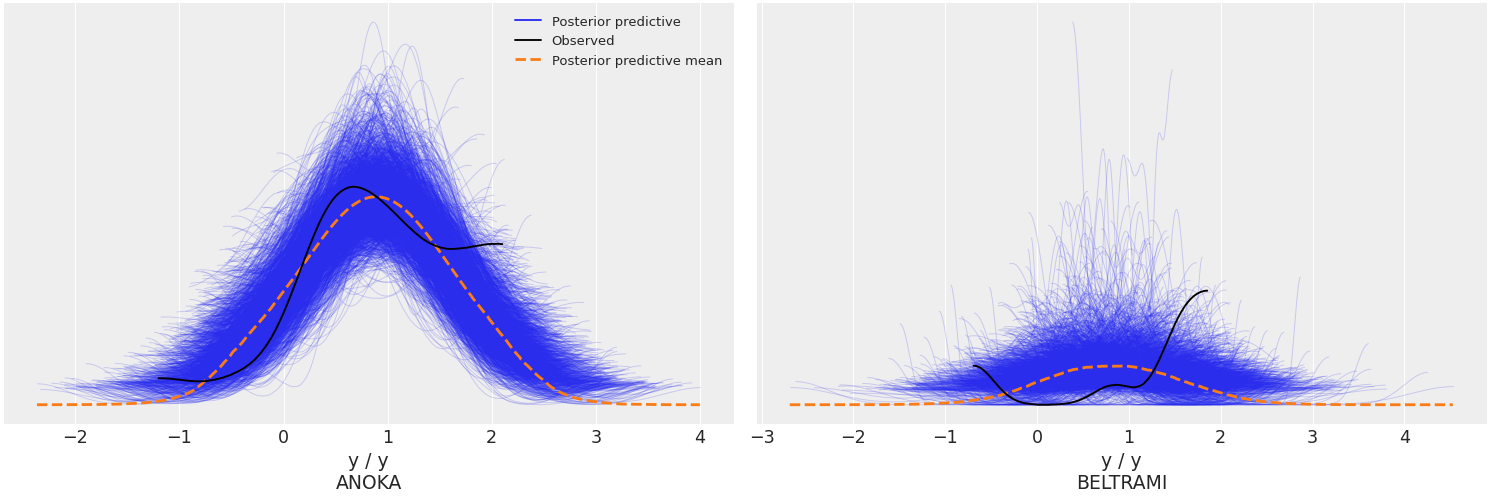
Use the
coordsandflattenparameters to plot selected variable dimensions across multiple plots. We will now modify the dimensionobs_idto contain indicate the name of the county where the measure was taken. The change has to be done on bothposterior_predictiveandobserved_datagroups, which is why we will usemap()to apply the same function to both groups. Afterwards, we will select the counties to be plotted with thecoordsarg.>>> obs_county = data.posterior["County"][data.constant_data["county_idx"]] >>> data = data.assign_coords(obs_id=obs_county, groups="observed_vars") >>> az.plot_ppc(data, coords={'obs_id': ['ANOKA', 'BELTRAMI']}, flatten=[])
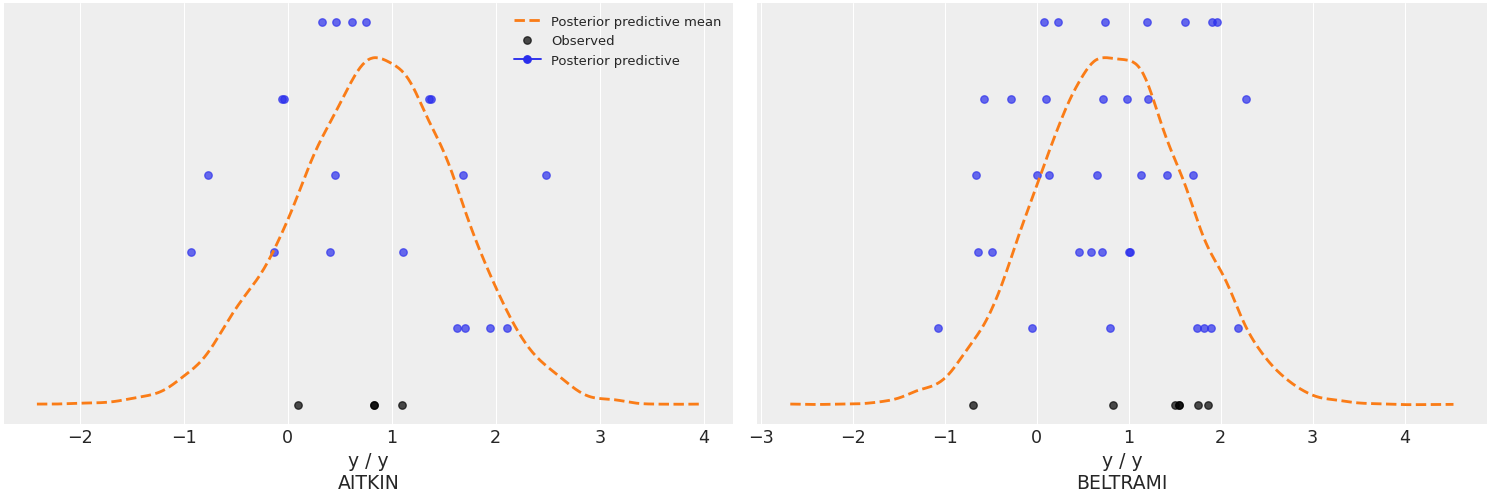
Plot the overlay using a stacked scatter plot that is particularly useful when the sample sizes are small.
>>> az.plot_ppc(data, kind='scatter', flatten=[], >>> coords={'obs_id': ['AITKIN', 'BELTRAMI']})
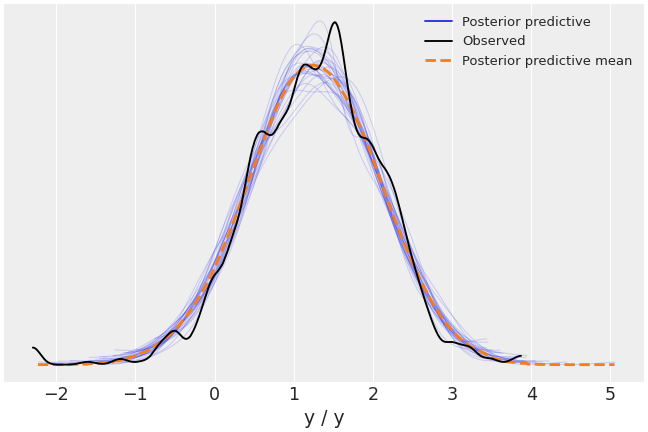
Plot random posterior predictive sub-samples.
>>> az.plot_ppc(data, num_pp_samples=30, random_seed=7)